Missing or censored data
Missing data
Neural networks do not naturally handle missing data, and this property can preclude their use in a broad range of applications. Here, we describe two techniques that alleviate this challenge in the context of parameter point estimation: the masking approach and the expectation-maximisation (EM) approach.
As a running example, we consider a Gaussian process model where the data are collected over a regular grid, but where some elements of the grid are unobserved. This situation often arises in, for example, remote-sensing applications, where the presence of cloud cover prevents measurement in some places. Below, we load the packages needed in this example, and define some aspects of the model that will remain constant throughout (e.g., the prior, the spatial domain). We also define types and functions for sampling from the prior distribution and for simulating marginally from the data model.
using NeuralEstimators, Flux
using Distributions: Uniform
using Distances, LinearAlgebra
using MLUtils: flatten
using SpecialFunctions: besselk, gamma
using Statistics: mean
# Prior and dimension of parameter vector
Π = (τ = Uniform(0, 1.0), ρ = Uniform(0, 0.4))
d = length(Π)
# Define the grid and compute the distance matrix
points = range(0, 1, 16)
S = expandgrid(points, points)
D = pairwise(Euclidean(), S, dims = 1)
# Collect model information for later use
ξ = (Π = Π, S = S, D = D)
# Struct for storing parameters and Cholesky factors
struct Parameters <: AbstractParameterSet
θ
L
end
# Matern covariance function
function matern(h, ρ, ν, σ² = one(typeof(h)))
@assert h >= 0 "h should be non-negative"
@assert ρ > 0 "ρ should be positive"
@assert ν > 0 "ν should be positive"
if h == 0
σ²
else
d = h / ρ
σ² * ((2^(1 - ν)) / gamma(ν)) * d^ν * besselk(ν, d)
end
end
# Constructor for above struct
function sample(K::Integer, ξ)
# Sample parameters from the prior
Π = ξ.Π
τ = rand(Π.τ, K)
ρ = rand(Π.ρ, K)
ν = 1 # fixed smoothness
# Compute Cholesky factors
L = map(1:K) do k
C = matern.(UpperTriangular(ξ.D), ρ[k], ν, 1)
L = cholesky(Symmetric(C)).L
convert(Array, L)
end
L = Base.stack(L)
# Concatenate into matrix
θ = permutedims(hcat(τ, ρ))
Parameters(θ, L)
end
# Marginal simulation from the data model
function simulate(parameters::Parameters, m::Integer)
K = size(parameters, 2)
τ = parameters.θ[1, :]
L = parameters.L
G = isqrt(size(L, 1)) # side-length of grid
Z = map(1:K) do k
z = simulategaussian(L[:, :, k], m)
z = z + τ[k] * randn(size(z)...)
z = reshape(z, G, G, 1, :)
z
end
return Z
endThe masking approach
The first missing-data technique that we consider is the so-called masking approach of Wang et al. (2024); see also the discussion by Sainsbury-Dale et al. (2025, Sec. 2.2). The strategy involves completing the data by replacing missing values with zeros, and using auxiliary variables to encode the missingness pattern, which are also passed into the network.
Let
where
The manner in which encodedata().
Since the missingness pattern removedata()):
# Marginal simulation from the data model and a MCAR missingness model
function simulatemissing(parameters::Parameters, m::Integer)
Z = simulate(parameters, m) # complete data
UW = map(Z) do z
prop = rand() # sample a missingness proportion
z = removedata(z, prop) # randomly remove a proportion of the data
uw = encodedata(z) # replace missing entries with zero and encode missingness pattern
uw
end
return UW
endNext, we construct and train a masked neural Bayes estimator using a CNN architecture. Here, the first convolutional layer takes two input channels, since we store the augmented data
# Construct DeepSet object
ψ = Chain(
Conv((10, 10), 2 => 16, relu),
Conv((5, 5), 16 => 32, relu),
Conv((3, 3), 32 => 64, relu),
flatten
)
ϕ = Chain(Dense(64, 256, relu), Dense(256, d, exp))
network = DeepSet(ψ, ϕ)
# Initialise point estimator
θ̂ = PointEstimator(network)
# Train the masked neural Bayes estimator
θ̂ = train(θ̂, sample, simulatemissing, simulator_args = 1, ξ = ξ, K = 1000, epochs = 10)Once trained, we can apply our masked neural Bayes estimator to (incomplete) observed data. The data must be encoded in the same manner as during training. Below, we use simulated data as a surrogate for real data, with a missingness proportion of 0.25:
θ = sample(1, ξ) # true parameters
Z = simulate(θ, 1)[1] # complete data
Z = removedata(Z, 0.25) # "observed" incomplete data (i.e., with missing values)
UW = encodedata(Z) # augmented data {U, W}
θ̂(UW) # point estimateThe EM approach
Let
where realisations of the missing-data component,
First, we construct a neural approximation of the MAP estimator. In this example, we will take tanhloss() in the limit
# Construct DeepSet object
ψ = Chain(
Conv((10, 10), 1 => 16, relu),
Conv((5, 5), 16 => 32, relu),
Conv((3, 3), 32 => 64, relu),
flatten
)
ϕ = Chain(
Dense(64, 256, relu),
Dense(256, d, exp)
)
network = DeepSet(ψ, ϕ)
# Initialise point estimator
θ̂ = PointEstimator(network)
# Train neural Bayes estimator
H = 50
θ̂ = train(θ̂, sample, simulate, simulator_args = H, ξ = ξ, K = 1000, epochs = 10)Next, we define a function for conditional simulation (see EM for details on the required format of this function):
function simulateconditional(Z::M, θ; nsims::Integer = 1, ξ) where {M <: AbstractMatrix{Union{Missing, T}}} where T
# Save the original dimensions
dims = size(Z)
# Convert to vector
Z = vec(Z)
# Compute the indices of the observed and missing data
I₁ = findall(z -> !ismissing(z), Z) # indices of observed data
I₂ = findall(z -> ismissing(z), Z) # indices of missing data
n₁ = length(I₁)
n₂ = length(I₂)
# Extract the observed data and drop Missing from the eltype of the container
Z₁ = Z[I₁]
Z₁ = [Z₁...]
# Distance matrices needed for covariance matrices
D = ξ.D # distance matrix for all locations in the grid
D₂₂ = D[I₂, I₂]
D₁₁ = D[I₁, I₁]
D₁₂ = D[I₁, I₂]
# Extract the parameters from θ
τ = θ[1]
ρ = θ[2]
# Compute covariance matrices
ν = 1 # fixed smoothness
Σ₂₂ = matern.(UpperTriangular(D₂₂), ρ, ν); Σ₂₂[diagind(Σ₂₂)] .+= τ^2
Σ₁₁ = matern.(UpperTriangular(D₁₁), ρ, ν); Σ₁₁[diagind(Σ₁₁)] .+= τ^2
Σ₁₂ = matern.(D₁₂, ρ, ν)
# Compute the Cholesky factor of Σ₁₁ and solve the lower triangular system
L₁₁ = cholesky(Symmetric(Σ₁₁)).L
x = L₁₁ \ Σ₁₂
# Conditional covariance matrix, cov(Z₂ ∣ Z₁, θ), and its Cholesky factor
Σ = Σ₂₂ - x'x
L = cholesky(Symmetric(Σ)).L
# Conditonal mean, E(Z₂ ∣ Z₁, θ)
y = L₁₁ \ Z₁
μ = x'y
# Simulate from the distribution Z₂ ∣ Z₁, θ ∼ N(μ, Σ)
z = randn(n₂, nsims)
Z₂ = μ .+ L * z
# Combine the observed and missing data to form the complete data
Z = map(1:nsims) do l
z = Vector{T}(undef, n₁ + n₂)
z[I₁] = Z₁
z[I₂] = Z₂[:, l]
z
end
Z = stack(Z)
# Convert Z to an array with appropriate dimensions
Z = reshape(Z, dims..., 1, nsims)
return Z
endNow we can use the neural EM algorithm to get parameter point estimates from data containing missing values. The algorithm is implemented with the type EM. Again, here we use simulated data as a surrogate for real data:
θ = sample(1, ξ) # true parameters
Z = simulate(θ, 1)[1][:, :] # complete data
Z = removedata(Z, 0.25) # "observed" incomplete data (i.e., with missing values)
θ₀ = mean.([Π...]) # initial estimate, the prior mean
neuralem = EM(simulateconditional, θ̂)
neuralem(Z, θ₀, nsims = H, ξ = ξ)Censored data
Neural estimators can be constructed to handle censored data as input, by exploiting the masking approach described above in the context of missing data. For simplicity, here we describe inference with left censored data (i.e., where we observe only those data that exceed some threshold), but extensions to right or interval censoring are possible. We first present the framework for General censoring, where data are considered censored based on an arbitrary, user-defined censoring scheme. We then consider Peaks-over-threshold censoring, a special case in which the data are treated as censored if they do not exceed their corresponding marginal
As a running example, we consider a bivariate random scale Gaussian mixture copula; see Engelke, Opitz, and Wadsworth (2019) and Huser and Wadsworth (2019). We consider the task of estimating
where
Simulation of the random scale mixture (on uniform margins) and its marginal ditribution function are provided below. Transforming the data to exponential margins can, in some cases, enhance training efficiency (Richards et al., 2024). However, for simplicity, we do not apply this transformation here.
# Libraries used throughout this example
using NeuralEstimators, Flux
using Folds
using CUDA # GPU if it is available
using LinearAlgebra: Symmetric, cholesky
using Distributions: cdf, Uniform, Normal, quantile
using CairoMakie
# Sampling θ from the prior distribution
function sample(K)
ρ = rand(Uniform(-0.99, 0.99), K)
δ = rand(Uniform(0.0, 1.0), K)
θ = vcat(ρ', δ')
return θ
end
# Marginal simulation of Z | θ
function simulate(θ, m)
Z = Folds.map(1:size(θ, 2)) do k
ρ = θ[1, k]
δ = θ[2, k]
Σ = [1 ρ; ρ 1]
L = cholesky(Symmetric(Σ)).L
X = L * randn(2, m) # Standard Gaussian margins
X = -log.(1 .- cdf.(Normal(), X)) # Transform to unit exponential margins
R = -log.(1 .- rand(1, m))
Y = δ .* R .+ (1 - δ) .* X
Z = F.(Y; δ = δ) # Transform to uniform margins
end
return Z
end
# Marginal distribution function; see Huser and Wadsworth (2019)
function F(y; δ)
if δ == 0.5
u = 1 .- exp.(- 2 .* y) .* (1 .+ 2 .* y)
else
u = 1 .- (δ ./ (2 .* δ .- 1)) .* exp.(- y ./ δ) .+ ((1 .- δ) ./ (2 * δ .- 1)) .* exp.( - y ./ (1 - δ))
end
return u
endGeneral censoring
Inference with censored data can proceed in an analogous manner to the The masking approach for missing data. First, consider a vector
where
The manner in which
The following helper function implements a simple version of the general censoring framework described above, based on a vector of censoring levels
# Constructing augmented data from Z and the censoring threshold c
function censorandaugment(Z; c, v = -1.0)
W = 1 * (Z .<= c)
U = ifelse.(Z .<= c, v, Z)
return vcat(U, W)
endThe above censoring function can then be incorporated into the data simulator as follows.
# Marginal simulation of censored data
function simulatecensored(θ, m; kwargs...)
Z = simulate(θ, m)
UW = Folds.map(Z) do Zₖ
mapslices(Z -> censorandaugment(Z; kwargs...), Zₖ, dims = 1)
end
endBelow, we construct a neural point estimator for censored data, based on a DeepSet architecture.
n = 2 # dimension of each data replicate (bivariate)
w = 128 # width of each hidden layer
# Final layer has output dimension d=2 and enforces parameter constraints
final_layer = Parallel(
vcat,
Dense(w, 1, tanh), # ρ ∈ (-1,1)
Dense(w, 1, sigmoid) # δ ∈ (0,1)
)
ψ = Chain(Dense(n * 2, w, relu), Dense(w, w, relu))
ϕ = Chain(Dense(w, w, relu), final_layer)
network = DeepSet(ψ, ϕ)
# Initialise the estimator
estimator = PointEstimator(network)We now train and assess two estimators for censored data; one with c = [0, 0] and one with c = [0.5, 0.5]. When the data c can be interpreted as the expected number of censored values in each component; thus, c = [0, 0] corresonds to no censoring of the data, and c = [0.5, 0.5] corresponds to a situation where, on average, 50% of each dimension
# Number of independent replicates in each data set
m = 200
# Train an estimator with no censoring
simulator1(θ, m) = simulatecensored(θ, m; c = [0, 0])
estimator1 = train(estimator, sample, simulator1, simulator_args = m)
# Train an estimator with mild censoring
simulator2(θ, m) = simulatecensored(θ, m; c = [0.5, 0.5])
estimator2 = train(estimator, sample, simulator2, simulator_args = m)
# Assessment
θ_test = sample(1000)
UW_test1 = simulator1(θ_test, m)
UW_test2 = simulator2(θ_test, m)
assessment1 = assess(estimator1, θ_test, UW_test1, parameter_names = ["ρ", "δ"], estimator_name = "No censoring")
assessment2 = assess(estimator2, θ_test, UW_test2, parameter_names = ["ρ", "δ"], estimator_name = "Mild censoring")
assessment = merge(assessment1, assessment2)
rmse(assessment)
plot(assessment)| Estimator | Parameter | RMSE |
|---|---|---|
| No censoring | ρ | 0.238167 |
| No censoring | δ | 0.100688 |
| Mild censoring | ρ | 0.394838 |
| Mild censoring | δ | 0.135169 |
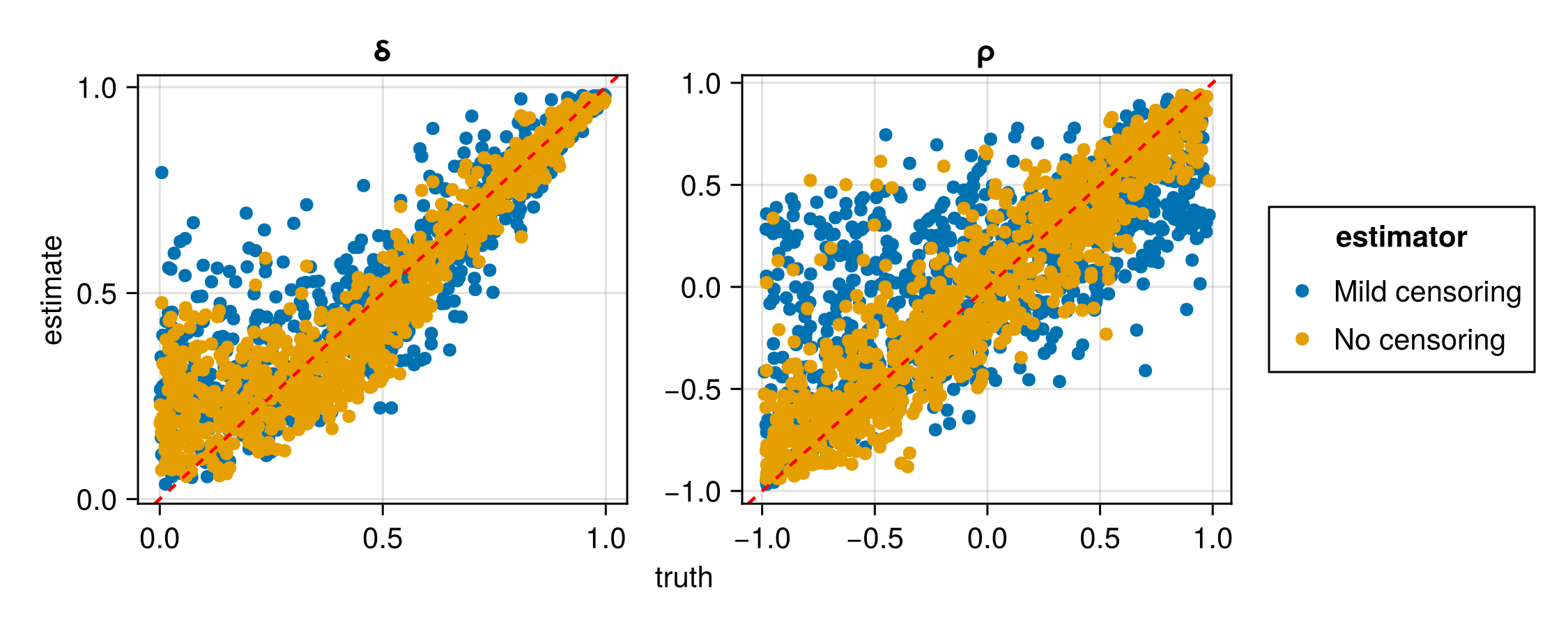
Here we have trained two separate neural estimators to handle two different censoring threshold vectors. However, one could train a single neural estimator that caters for a range of censoring thresholds, c, by allowing it to vary with the data samples and using it as an input to the neural network. In the next section, we illustrate this in the context of peaks-over-threshold modelling, whereby a single censoring threshold is defined to be the marginal c.
Peaks-over-threshold censoring
Richards et al. (2024) discuss neural Bayes estimation from censored data in the context of peaks-over-threshold extremal dependence modelling, where deliberate censoring of data is imposed to reduce estimation bias in the presence of marginally non-extreme events. In these settings, data are treated as censored if they do not exceed their corresponding marginal
Peaks-over-threshold censoring, with
Below, we sample a fixed set of
# Sampling values of τ to use during training
function sampleτ(K)
τ = rand(Uniform(0.0, 0.9), 1, K)
end
# Adapt the censored data simulation to allow for τ as an input
function simulatecensored(θ, τ, m; kwargs...)
Z = simulate(θ, m)
K = size(θ, 2)
UW = Folds.map(1:K) do k
Zₖ = Z[k]
τₖ = τ[k]
cₖ = τₖ # data are on uniform margins: censoring threshold equals τ
mapslices(Z -> censorandaugment(Z; c = cₖ, kwargs...), Zₖ, dims = 1)
end
end
# Generate the data used for training and validation
K = 50000 # number of training samples
m = 500 # number of replicates in each data set
θ_train = sample(K)
θ_val = sample(K ÷ 5)
τ_train = sampleτ(K)
τ_val = sampleτ(K ÷ 5)
UW_train = simulatecensored(θ_train, τ_train, m)
UW_val = simulatecensored(θ_val, τ_val, m)In this example, the probability level DeepSet architecture. To do this, we increase the input dimension of the outer network by one, and then combine the data DeepSet for details).
# Construct neural network based on DeepSet architecture
ψ = Chain(Dense(n * 2, w, relu),Dense(w, w, relu))
ϕ = Chain(Dense(w + 1, w, relu), final_layer)
network = DeepSet(ψ, ϕ)
# Initialise the estimator
estimator = PointEstimator(network)
# Train the estimator
estimator = train(estimator, θ_train, θ_val, (UW_train, τ_train), (UW_val, τ_val))Our trained estimator can now be used for any value of
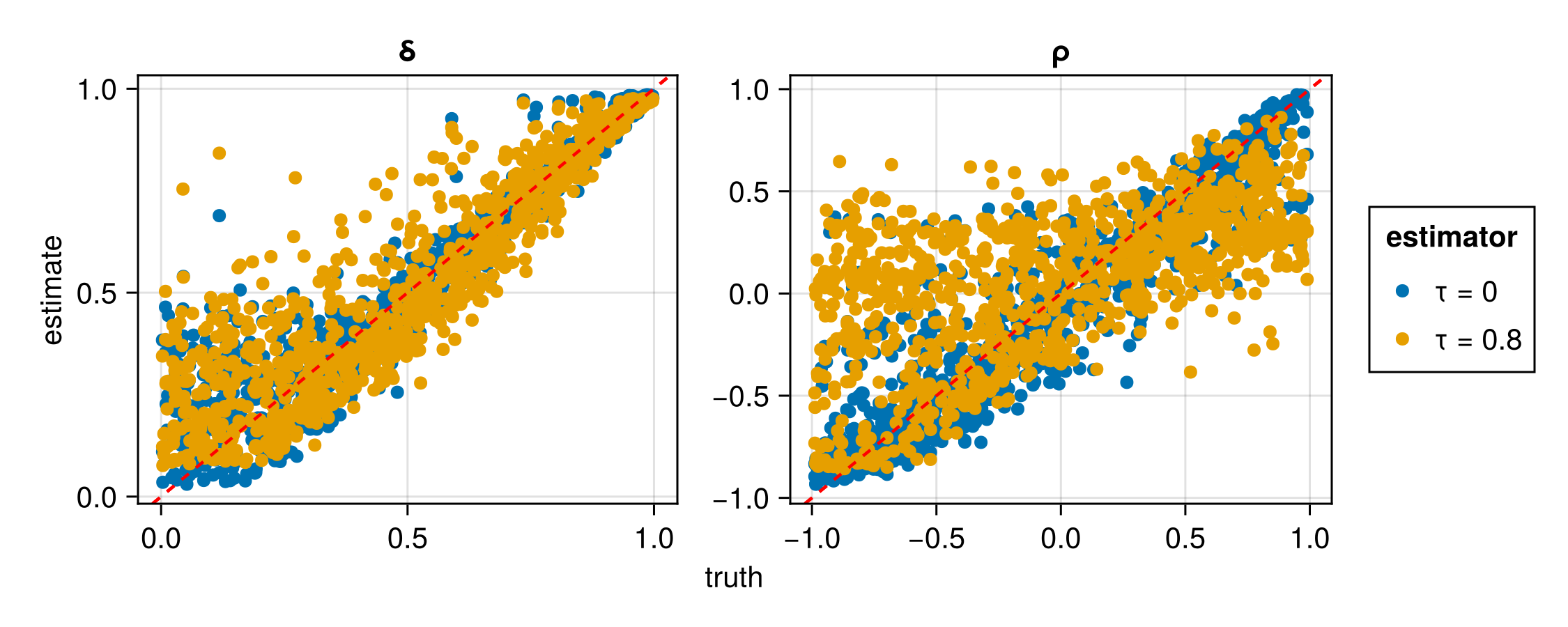
Below, we assess the estimator for different values of
# Test parameters
θ_test = sample(1000)
# Assessment with τ fixed to 0 (no censoring)
τ_test1 = fill(0.0, 1000)'
UW_test1 = simulatecensored(θ_test, τ_test1, m)
assessment1 = assess(estimator, θ_test, (UW_test1, τ_test1), parameter_names = ["ρ", "δ"], estimator_name = "τ = 0")
# Assessment with τ fixed to 0.8
τ_test2 = fill(0.8, 1000)'
UW_test2 = simulatecensored(θ_test, τ_test2, m)
assessment2 = assess(estimator, θ_test, (UW_test2, τ_test2), parameter_names = ["ρ", "δ"], estimator_name = "τ = 0.8")
# Compare results between the two censoring probability levels
assessment = merge(assessment1, assessment2)
rmse(assessment)
plot(assessment)| Estimator | Parameter | RMSE |
|---|---|---|
| τ = 0 | ρ | 0.241780 |
| τ = 0 | δ | 0.096779 |
| τ = 0.80 | ρ | 0.476348 |
| τ = 0.80 | δ | 0.124913 |