Spatio-temporal data
Here, we develop a neural estimator for data modelled as a Spatio-Temporal Gaussian Random Field (GRF). This model can be used for environmental phenomena such as sea surface temperature (SST) that exhibit correlation across both space and time.
We assume that the data are observed at locations
where
where
We place independent uniform priors on the parameters:
Package dependencies
using NeuralEstimators
using NeuralEstimators: getobs, numobs
using Flux
using Folds # parallel simulation (start Julia with --threads=auto)
using Distances
using Distributions: Uniform
using LinearAlgebra
using Statistics
using CairoMakieThis example uses convolutional neural networks (CNNs), which are computationally intensive but highly parallelisable and therefore greatly benefit from GPU acceleration. Various GPU backends are supported and, once the relevant package is loaded and a compatible GPU is available, it will be used automatically:
using CUDA, cuDNNusing AMDGPUusing Metalusing oneAPISampling parameters
We first define a function to sample parameters from the prior distribution. Here, we store the parameters as a NamedMatrix so that parameter estimates are automatically labelled, though this is not required:
function sampler(K)
NamedMatrix(
θ₁ = rand(Uniform(0.05, 0.5), K),
θ₂ = rand(Uniform(0.1, 0.9), K)
)
endSimulating data and constructing the neural network
The simulator and the neural network are designed in tandem: the architecture is chosen based on the structure of the data, and the simulator must return data in a format appropriate for the chosen architecture.
The data are spatio-temporally dependent and observed over a regular spatio-temporal grid. Several architectures could handle these data correctly:
CNN with
channel dimensions: treats each time step as a channel. Simple and effective but requires fixed . CNN + RNN/LSTM/Transformer: a CNN extracts spatial features from each time slice, and an RNN/LSTM/Transformer processes them sequentially to capture temporal dependence. Handles variable
naturally but more computationally expensive. DeepSet with CNN inner network: for an AR(1) process, the first differences
DeepSetwith a CNN inner network applied to each difference field can be used. Handles variablenaturally and computationally efficiently.
We adopt the last approach, as it is simple to implement, exploits the model structure, and is computationally efficient.
DeepSet objects act on vectors, where each element of the vector is associated with one parameter vector
# Matérn covariance with unknown range parameter and fixed smoothness parameter 3/2
function matern15(d, ρ)
d == 0.0 && return 1.0
r = sqrt(3) * d / ρ
return (1 + r) * exp(-r)
end
# Compute Δₜ for t = 2, ..., T
function firstdifference(Z)
# Equivalently: Z[:, :, :, 2:end] - Z[:, :, :, 1:end-1]
getobs(Z, 2:numobs(Z)) - getobs(Z, 1:numobs(Z)-1)
end
# Simulate a spatio-temporal field with T time steps
function simulator(θ::AbstractVector, T::Integer; grid_size = 10, first_difference::Bool = true)
ρ = θ["θ₁"]
φ = θ["θ₂"]
# Regular spatial grid
locs = [(i/grid_size, j/grid_size) for i in 1:grid_size for j in 1:grid_size]
n_spatial = length(locs)
# Spatial covariance matrix and its Cholesky factor
Σ = [matern15(norm(collect(locs[i]) .- collect(locs[j])), ρ) for i in 1:n_spatial, j in 1:n_spatial]
L = cholesky(Σ + 1e-6 * I).L
# Dynamic system
Z = zeros(n_spatial, T)
Z[:, 1] = L * randn(n_spatial)
for t in 2:T
Z[:, t] = φ .* Z[:, t-1] .+ sqrt(1 - φ^2) .* (L * randn(n_spatial))
end
# Reshape into format required by our chosen architecture
Z = reshape(Z, grid_size, grid_size, 1, T)
return first_difference ? firstdifference(Z) : Z
end
simulator(θ::AbstractVector, T; kwargs...) = simulator(θ, rand(T); kwargs...)
simulator(θ::AbstractMatrix, T = 10:30; kwargs...) = Folds.map(ϑ -> simulator(ϑ, T; kwargs...), eachcol(θ))With this data format, the DeepSet inner network ψ processes each difference field ϕ maps the aggregated latent features to summary statistics:
d = 2 # number of parameters
w = 128 # hidden layer width
nc = 32 # channels in final conv layer and dimension of DeepSet latent space
num_summaries = 3d # number of summaries output by the network
ψ = Chain(
Conv((3, 3), 1 => 16, relu; pad = 1),
MaxPool((2, 2)),
Conv((3, 3), 16 => nc, relu; pad = 1),
GlobalMeanPool(),
Flux.flatten,
)
ϕ = Chain(
Dense(nc, w, relu),
Dense(w, w, relu),
Dense(w, w, relu),
Dense(w, num_summaries),
)
network = DeepSet(ψ, ϕ)Bonus: Visualizing spatio-temporal dependence
Before initializing the neural estimator with our chosen neural-network architecture, let's simulate and visualize some realizations from the model.
We'll focus on the effect of the temporal range parameter,
# Fix θ₁, vary θ₂
θ = NamedMatrix(
θ₁ = [0.3, 0.3],
θ₂ = [0.1, 0.8]
)
# Simulate several time steps
Z = simulator(θ, 5; grid_size = 32, first_difference = false)
labels = [
"Low temporal dependence (θ₂ = 0.1)",
"High temporal dependence (θ₂ = 0.8)"
]
fig = Figure(size = (1350, 600));
for k in 1:2
Z_k = Z[k]
T = size(Z_k, 4)
for t in 1:T
ax = Axis(
fig[k, t],
title = t == 1 ? labels[k] : "t = $t",
aspect = DataAspect()
)
hidedecorations!(ax)
hm = heatmap!(
ax,
Z_k[:, :, 1, t],
colormap = :balance,
colorrange = (-2.5, 2.5)
)
if t == T
Colorbar(fig[k, T + 1], hm)
end
end
end
display(fig)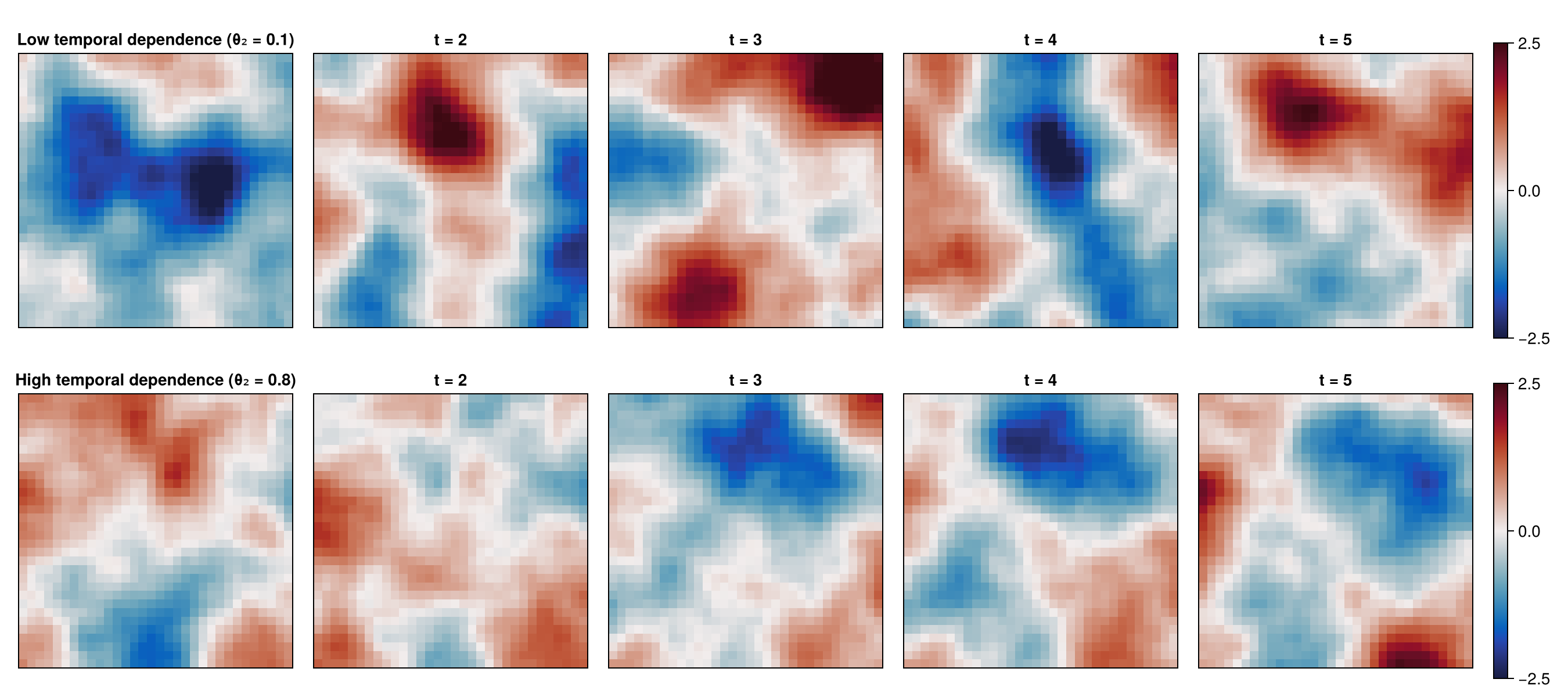
Constructing the neural estimator
We now construct a neural estimator by wrapping the neural network in the subtype corresponding to the intended inferential method:
estimator = PointEstimator(network, d; num_summaries = num_summaries)estimator = PosteriorEstimator(network, d; num_summaries = num_summaries)estimator = RatioEstimator(network, d; num_summaries = num_summaries)Training the estimator
Next, we train the estimator using train. One may pass sampler and simulator directly to train for on-the-fly simulation, but here we pre-simulate fixed training sets, which can be faster when the simulator is expensive:
K = 2500 # size of the training set
θ_train = sampler(K)
θ_val = sampler(K)
Z_train = simulator(θ_train);
Z_val = simulator(θ_val);
estimator = train(estimator, θ_train, θ_val, Z_train, Z_val)
# Alternatively, simulate on-the-fly:
# estimator = train(estimator, sampler, simulator; K = K)The empirical risk (average loss) over the training and validation sets can be plotted using plotrisk.
One may wish to save a trained estimator and load it in a later session: see Saving and loading estimators for details on how this can be done.
Assessing the estimator
The function assess can be used to assess the trained estimator:
θ_test = sampler(1000)
Z_test = simulator(θ_test, 20);
assessment = assess(estimator, θ_test, Z_test)The resulting Assessment object contains ground-truth parameters, estimates, and other quantities that can be used to compute quantitative and qualitative diagnostics:
bias(assessment) # θ₁ = ..., θ₂ = ...
rmse(assessment) # θ₁ = ..., θ₂ = ...
plot(assessment)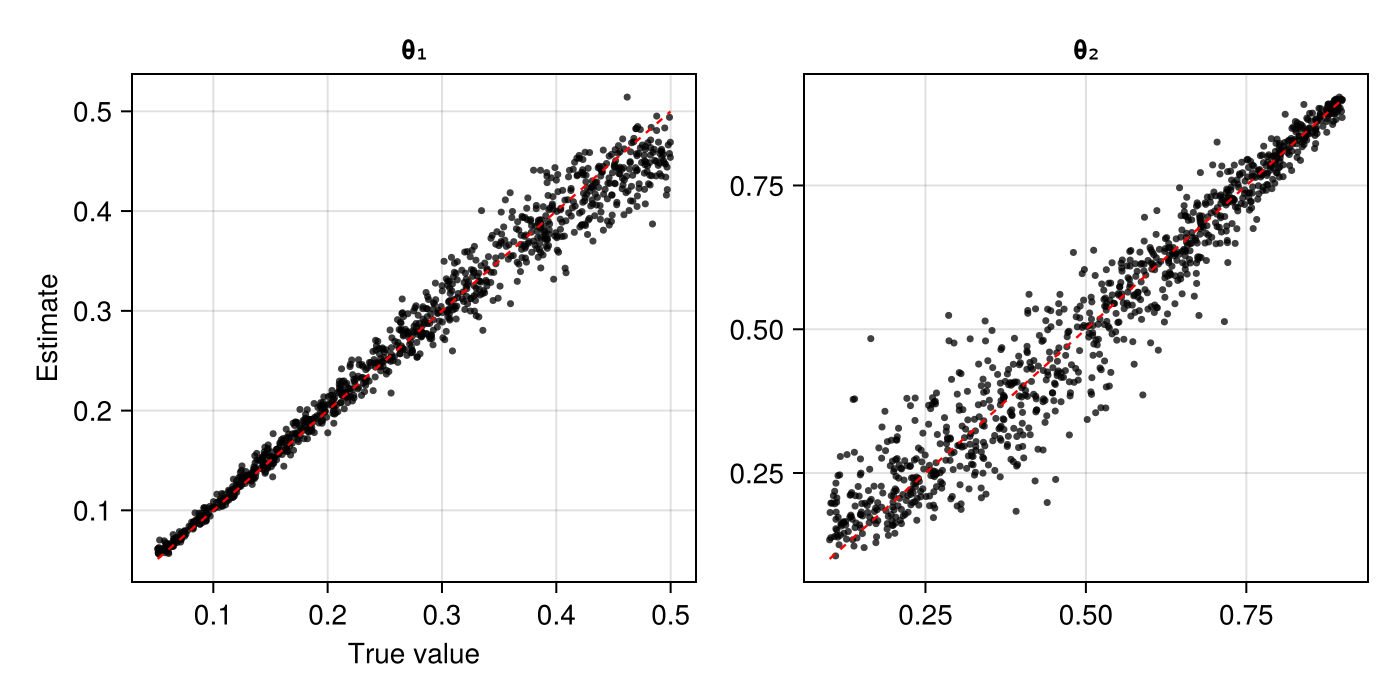
Applying the estimator to observed data
Once an estimator is deemed to be well calibrated, it may be applied to observed data (below, we use simulated data as a stand-in for observed data):
θ = sampler(1)
Z = simulator(θ, 20);estimate(estimator, Z)sampleposterior(estimator, Z)sampleposterior(estimator, Z)