Exchangeable data
Here, we develop a neural estimator to infer
Package dependencies
using NeuralEstimators
using Flux
using Distributions: InverseGamma
using CairoMakieTo improve computational efficiency, various GPU backends are supported. Once the relevant package is loaded and a compatible GPU is available, it will be used automatically:
using CUDA, cuDNNusing AMDGPUusing Metalusing oneAPISampling parameters
We first define a function to sample parameters from the prior distribution. Here, we store the parameters as a NamedMatrix so that parameter estimates are automatically labelled, though this is not required:
function sampler(K)
NamedMatrix(
μ = randn(K),
σ = rand(InverseGamma(3, 1), K)
)
endSimulating data
Next, we define the statistical model implicitly through simulation. Our data consist of Vector. In this example each replicate Matrix with one row and
We define three methods for simulating data, illustrating Julia's multiple dispatch and short-form function syntax: the first simulates a data set of fixed size
simulator(θ::AbstractVector, m::Integer) = θ["μ"] .+ θ["σ"] .* randn(1, m)
simulator(θ::AbstractVector, m) = simulator(θ, rand(m))
simulator(θ::AbstractMatrix, m = 10:100) = [simulator(ϑ, m) for ϑ in eachcol(θ)]Constructing the neural network
In this package, the neural network specified by the user is typically a summary network that transforms data into a vector of
In this example, our data are replicated, and we therefore adopt the DeepSets framework, implemented via DeepSet. A DeepSet consists of three components: an inner network that acts directly on each data replicate; a function that aggregates the outputs of the inner network; and an outer network (typically an MLP) that maps the aggregated output to
n = 1 # dimension of each data replicate (univariate)
d = 2 # dimension of the parameter vector θ
num_summaries = 3d # number of summary statistics for θ
w = 64 # width of each hidden layer
ψ = Chain(Dense(n, w, relu), Dense(w, w, relu)) # inner network
ϕ = Chain(Dense(w, w, relu), Dense(w, num_summaries)) # outer network
network = DeepSet(ψ, ϕ)Constructing the neural estimator
We now construct a neural estimator by wrapping the neural network in the subtype corresponding to the intended inferential method:
estimator = PointEstimator(network, d; num_summaries = num_summaries)estimator = PosteriorEstimator(network, d; num_summaries = num_summaries)estimator = RatioEstimator(network, d; num_summaries = num_summaries)Training the estimator
Next, we train the estimator using train. Below, we pass our user-defined functions for sampling parameters and simulating data, but one may also pass fixed parameter and/or data instances:
estimator = train(estimator, sampler, simulator)The empirical risk (average loss) over the training and validation sets can be plotted using plotrisk.
One may wish to save a trained estimator and load it in a later session: see Saving and loading estimators for details on how this can be done.
Assessing the estimator
The function assess can then be used to assess the trained estimator based on unseen test data simulated from the statistical model:
θ_test = sampler(1000) # test parameters
Z_test = simulator(θ_test, 50) # test data
assessment = assess(estimator, θ_test, Z_test)The resulting Assessment object contains ground-truth parameters, estimates, and other quantities that can be used to compute quantitative and qualitative diagnostics:
bias(assessment) # μ = 0.002, σ = 0.017
rmse(assessment) # μ = 0.086, σ = 0.078
risk(assessment) # 0.055
plot(assessment)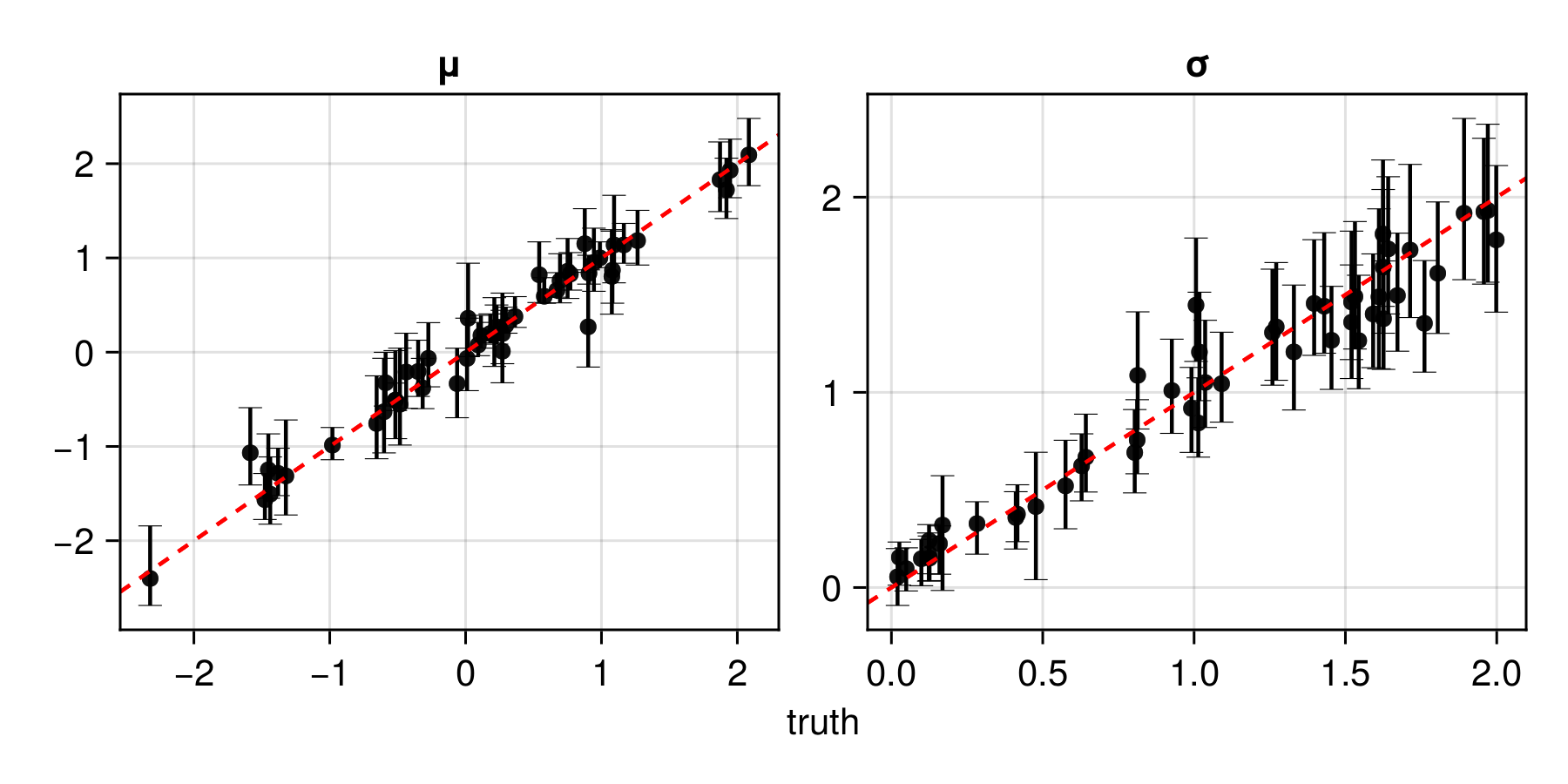
Applying the estimator to observed data
Once an estimator is deemed to be well calibrated, it may be applied to observed data (below, we use simulated data as a stand-in for observed data):
θ = sampler(1) # ground truth (not known in practice)
Z = simulator(θ, 100) # stand-in for real dataestimate(estimator, Z) # point estimate
interval(bootstrap(estimator, Z)) # 95% non-parametric bootstrap intervalssampleposterior(estimator, Z) # posterior samplesampleposterior(estimator, Z) # posterior sample